Digitizing Tissue Microarrays: Optimizing TMA Workflow with Whole Slide Imaging
Background:
Tissue microarray (TMA) technology has evolved from a high-throughput immunohistochemical screening tool into a central platform for spatial biology, multiplexed molecular profiling, and AI-enabled digital pathology. However, traditional TMA construction remains labor-intensive, susceptible to sampling variability, and limited in scalability. Concurrently, advanced multiplex and spatial assays impose significant financial and computational burdens, restricting large-cohort implementation. We developed an integrated, automated TMA framework designed to enhance reproducibility, scalability, and cost-efficiency in translational research and diagnostic workflows.
Design:
The integrated platform combines high-resolution whole-slide digitization using MoticEasyScan, AI-assisted region-of-interest (ROI) selection, and automated tissue array construction using a computer-driven TMA system (TMAtic). Digital slide overlays, barcode traceability, and metadata-linked outputs ensure reproducible array geometry and donor-to-core traceability. The system supports compatibility with conventional histology, multiplex immunohistochemistry, RNA in situ hybridization, spatial transcriptomics, and genomic core retrieval workflows. MoticEasyScan functions as the digital backbone of the workflow, enabling standardized image acquisition, scalable data management, and generation of AI-ready datasets before and after array construction.
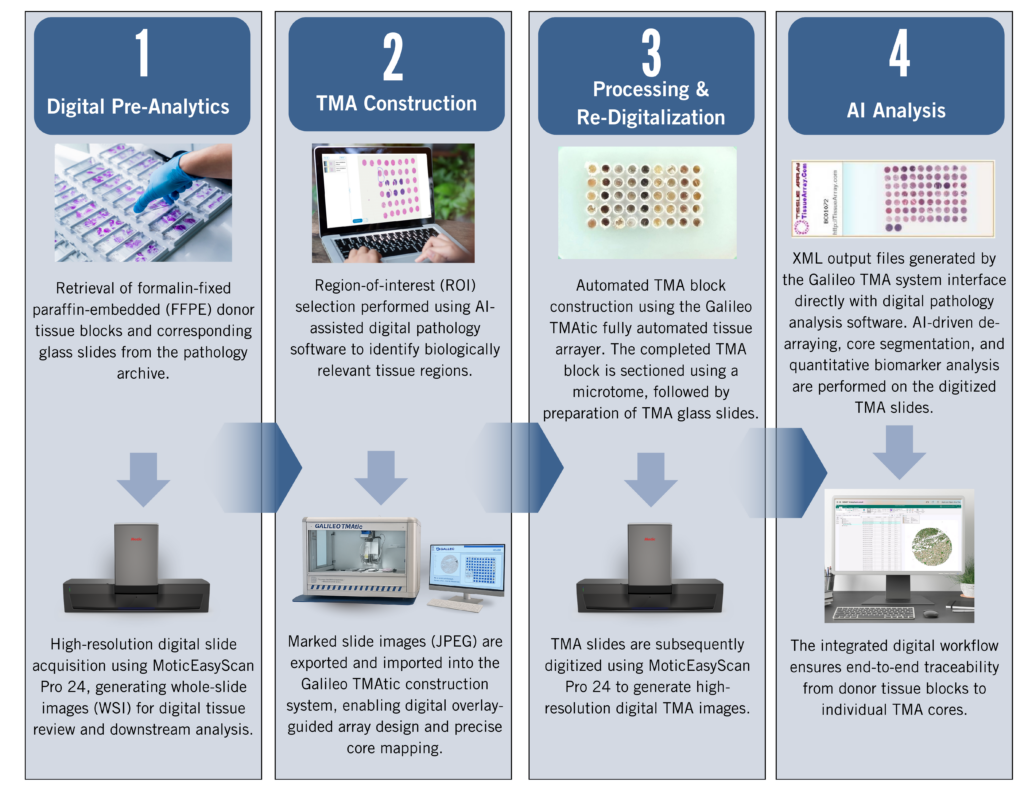
Figure 1: End-to-End Digital Tissue Microarray Workflow: From Digital Pre-Analytics and Automated TMA Construction to Re-Digitization and AI-Driven Analysis
Workflow:
The standardized TMA workflow begins with retrieval of formalin-fixed paraffin-embedded donor blocks and corresponding glass slides from the pathology archive. Prepared H&E, IHC, or multiplex-stained slides are then digitized at high resolution using MoticEasyScan, establishing a precise digital representation of the tissue architecture. Regions of interest are selected using AI-assisted digital pathology and spatial analysis tools, allowing targeted identification of biologically relevant areas such as tumor margins, immune niches, or stromal compartments. Based on these annotated regions, tissue cores are physically transferred into recipient blocks using a manual, semi-automated, or fully automated arraying instrument, with computer-driven systems enabling digital slide overlay guidance and standardized array geometry. Following construction, completed TMA slides are rescanned using MoticEasyScan to generate high-resolution images for downstream analysis. Finally, AI-enabled digital pathology tools perform automated de-arraying, single-core annotation, and quantitative biomarker assessment, ensuring reproducible core-level analysis and complete donor-to-core traceability.
Results:
Digital pre-selection enhances sampling precision and reduces operator-dependent variability. Automated construction improves alignment consistency and throughput across large cohorts. Consolidation of dozens to hundreds of specimens onto a single slide reduces per-sample costs for IHC and multiplex assays by up to 70%, with amplified savings in spatial biology applications. TMA-focused imaging further reduces digital storage requirements and computational demand compared to whole-section spatial assays, supporting scalable AI-based analysis in multicenter and biobank-driven studies.
Conclusion:
In the era of AI and spatial biology, high-throughput TMAs supported by integrated digital pathology platforms such as MoticEasyScan provide a scalable, cost-efficient infrastructure for research and diagnostic innovation. The convergence of automation, digital scanning, and AI-driven analysis transforms TMAs into reproducible, programmable platforms aligned with the evolving demands of precision pathology.